PIAS 1 Polyclonal Antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB, IHC-P |
|---|---|
| Primary Accession | O75925 |
| Reactivity | Human, Mouse |
| Host | Rabbit |
| Clonality | Polyclonal |
| Calculated MW | 71836 Da |
| Gene ID | 8554 |
|---|---|
| Other Names | PIAS1; DDXBP1; E3 SUMO-protein ligase PIAS1; DEAD/H box-binding protein 1; Gu-binding protein; GBP; Protein inhibitor of activated STAT protein 1; RNA helicase II-binding protein |
| Dilution | WB~~Western Blot: 1/500 - 1/2000. Immunohistochemistry: 1/100 - 1/300. ELISA: 1/10000. Not yet tested in other applications. |
| Format | Liquid in PBS containing 50% glycerol, 0.5% BSA and 0.09% (W/V) sodium azide. |
| Storage Conditions | -20℃ |
| Name | PIAS1 |
|---|---|
| Synonyms | DDXBP1 |
| Function | Functions as an E3-type small ubiquitin-like modifier (SUMO) ligase, stabilizing the interaction between UBE2I and the substrate, and as a SUMO-tethering factor (PubMed:11583632, PubMed:11867732, PubMed:14500712, PubMed:21965678, PubMed:36050397). Catalyzes sumoylation of various proteins, such as CEBPB, MRE11, MTA1, PTK2 and PML (PubMed:11583632, PubMed:11867732, PubMed:14500712, PubMed:21965678, PubMed:36050397). Plays a crucial role as a transcriptional coregulation in various cellular pathways, including the STAT pathway, the p53 pathway and the steroid hormone signaling pathway (PubMed:11583632, PubMed:11867732). In vitro, binds A/T-rich DNA (PubMed:15133049). The effects of this transcriptional coregulation, transactivation or silencing, may vary depending upon the biological context (PubMed:11583632, PubMed:11867732, PubMed:14500712, PubMed:21965678, PubMed:36050397). Mediates sumoylation of MRE11, stabilizing MRE11 on chromatin during end resection (PubMed:36050397). Sumoylates PML (at 'Lys-65' and 'Lys-160') and PML-RAR and promotes their ubiquitin-mediated degradation (By similarity). PIAS1-mediated sumoylation of PML promotes its interaction with CSNK2A1/CK2 which in turn promotes PML phosphorylation and degradation (By similarity). Enhances the sumoylation of MTA1 and may participate in its paralog- selective sumoylation (PubMed:21965678). Plays a dynamic role in adipogenesis by promoting the SUMOylation and degradation of CEBPB (By similarity). Mediates the nuclear mobility and localization of MSX1 to the nuclear periphery, whereby MSX1 is brought into the proximity of target myoblast differentiation factor genes (By similarity). Also required for the binding of MSX1 to the core enhancer region in target gene promoter regions, independent of its sumoylation activity (By similarity). Capable of binding to the core enhancer region TAAT box in the MYOD1 gene promoter (By similarity). |
| Cellular Location | Nucleus {ECO:0000250|UniProtKB:O88907}. Nucleus speckle Nucleus, PML body {ECO:0000250|UniProtKB:O88907}. Cytoplasm, cytoskeleton. Note=Interaction with CSRP2 may induce a partial redistribution along the cytoskeleton (PubMed:11672422). Interaction with MSX1 is required for localization to the nuclear periphery (By similarity) {ECO:0000250|UniProtKB:O88907, ECO:0000269|PubMed:11672422} |
| Tissue Location | Expressed in numerous tissues with highest level in testis. |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
Functions as an E3-type small ubiquitin-like modifier (SUMO) ligase, stabilizing the interaction between UBE2I and the substrate, and as a SUMO-tethering factor. Plays a crucial role as a transcriptional coregulation in various cellular pathways, including the STAT pathway, the p53 pathway and the steroid hormone signaling pathway. In vitro, binds A/T-rich DNA. The effects of this transcriptional coregulation, transactivation or silencing, may vary depending upon the biological context. Sumoylates PML (at'Lys-65' and 'Lys-160') and PML-RAR and promotes their ubiquitin-mediated degradation. PIAS1-mediated sumoylation of PML promotes its interaction with CSNK2A1/CK2 which in turn promotes PML phosphorylation and degradation (By similarity). Enhances the sumoylation of MTA1 and may participate in its paralog-selective sumoylation. Plays a dynamic role in adipogenesis by promoting the SUMOylation and degradation of CEBPB (By similarity).
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.




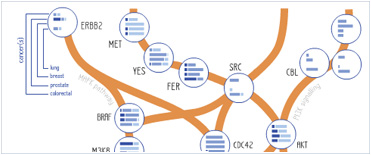







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.


