COP1 Polyclonal Antibody
- SPECIFICATION
- CITATIONS
- PROTOCOLS
- BACKGROUND

Application
| WB |
|---|---|
| Primary Accession | Q8NHY2 |
| Reactivity | Human, Mouse |
| Host | Rabbit |
| Clonality | Polyclonal |
| Calculated MW | 80474 Da |
| Gene ID | 64326 |
|---|---|
| Other Names | RFWD2; COP1; RNF200; E3 ubiquitin-protein ligase RFWD2; Constitutive photomorphogenesis protein 1 homolog; hCOP1; RING finger and WD repeat domain protein 2; RING finger protein 200 |
| Dilution | WB~~Western Blot: 1/500 - 1/2000. ELISA: 1/40000. Not yet tested in other applications. |
| Format | Liquid in PBS containing 50% glycerol, 0.5% BSA and 0.09% (W/V) sodium azide. |
| Storage Conditions | -20℃ |
| Name | COP1 (HGNC:17440) |
|---|---|
| Function | E3 ubiquitin-protein ligase that mediates ubiquitination and subsequent proteasomal degradation of target proteins. E3 ubiquitin ligases accept ubiquitin from an E2 ubiquitin-conjugating enzyme in the form of a thioester and then directly transfers the ubiquitin to targeted substrates. Involved in JUN ubiquitination and degradation. Directly involved in p53 (TP53) ubiquitination and degradation, thereby abolishing p53-dependent transcription and apoptosis. Ubiquitinates p53 independently of MDM2 or RCHY1. Probably mediates E3 ubiquitin ligase activity by functioning as the essential RING domain subunit of larger E3 complexes. In contrast, it does not constitute the catalytic RING subunit in the DCX DET1-COP1 complex that negatively regulates JUN, the ubiquitin ligase activity being mediated by RBX1. Involved in 14-3-3 protein sigma/SFN ubiquitination and proteasomal degradation, leading to AKT activation and promotion of cell survival. Ubiquitinates MTA1 leading to its proteasomal degradation. Upon binding to TRIB1, ubiquitinates CEBPA, which lacks a canonical COP1-binding motif (Probable). |
| Cellular Location | Nucleus speckle. Cytoplasm. Note=In the nucleus, it forms nuclear speckles |
| Tissue Location | Ubiquitously expressed at low level. Expressed at higher level in testis, placenta, skeletal muscle and heart |

Thousands of laboratories across the world have published research that depended on the performance of antibodies from Abcepta to advance their research. Check out links to articles that cite our products in major peer-reviewed journals, organized by research category.
info@abcepta.com, and receive a free "I Love Antibodies" mug.
Provided below are standard protocols that you may find useful for product applications.
Background
E3 ubiquitin-protein ligase that mediates ubiquitination and subsequent proteasomal degradation of target proteins. E3 ubiquitin ligases accept ubiquitin from an E2 ubiquitin- conjugating enzyme in the form of a thioester and then directly transfers the ubiquitin to targeted substrates. Involved in JUN ubiquitination and degradation. Directly involved in p53 (TP53) ubiquitination and degradation, thereby abolishing p53-dependent transcription and apoptosis. Ubiquitinates p53 independently of MDM2 or RCHY1. Probably mediates E3 ubiquitin ligase activity by functioning as the essential RING domain subunit of larger E3 complexes. In contrast, it does not constitute the catalytic RING subunit in the DCX DET1-COP1 complex that negatively regulates JUN, the ubiquitin ligase activity being mediated by RBX1. Involved in 14-3-3 protein sigma/SFN ubiquitination and proteasomal degradation, leading to AKT activation and promotion of cell survival. Ubiquitinates MTA1 leading to its proteasomal degradation. Upon binding to TRIB1, ubiquitinates CEBPA, which lacks a canonical COP1-binding motif (Probable).
If you have used an Abcepta product and would like to share how it has performed, please click on the "Submit Review" button and provide the requested information. Our staff will examine and post your review and contact you if needed.
If you have any additional inquiries please email technical services at tech@abcepta.com.




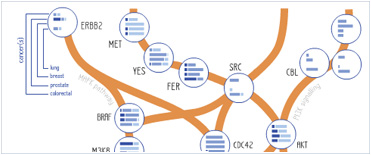







 Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them.
Foundational characteristics of cancer include proliferation, angiogenesis, migration, evasion of apoptosis, and cellular immortality. Find key markers for these cellular processes and antibodies to detect them. The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle.
The SUMOplot™ Analysis Program predicts and scores sumoylation sites in your protein. SUMOylation is a post-translational modification involved in various cellular processes, such as nuclear-cytosolic transport, transcriptional regulation, apoptosis, protein stability, response to stress, and progression through the cell cycle. The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.
The Autophagy Receptor Motif Plotter predicts and scores autophagy receptor binding sites in your protein. Identifying proteins connected to this pathway is critical to understanding the role of autophagy in physiological as well as pathological processes such as development, differentiation, neurodegenerative diseases, stress, infection, and cancer.


